Introduction
Invasive species represent a pervasive and formidable consequence of globalization, resulting in significant economic damage and posing substantial threats to indigenous biodiversity (Gethöffer and Siebert, Reference Gethöffer and Siebert2020). Through their disruptive effects on ecosystems, these species cause declines in biodiversity and may precipitate species extinctions via direct predatory activities (Atkinson, Reference Atkinson1996). Additionally, they frequently act as vectors for zoonotic diseases or serve as reservoir hosts for such pathogens.
Myocastor coypus, commonly known as the nutria or coypu, is prominently featured among the 100 most detrimental invasive species worldwide (Lowe et al., Reference Lowe, Browne, Boudjelas and De Poorter2000) and is formally recognized within the Catalogue of Invasive Alien Species of the European Union (European Commission, 2022). This semi-aquatic rodent, native to South America, is currently distributed across all continents except Australia (and New Zealand) and Antarctica (Woods et al., Reference Woods, Contreras, Willner-Chapman and Whidden1992).
The introduction of nutria to the Czech Republic during the early 20th century, primarily driven by fur trading (Anděra, Reference Anděra2011), typifies its successful establishment beyond its native range. Currently, nutrias occupy more than two-thirds of the country's territory (AOPK ČR, 2023). Although there is no general overpopulation, in urban areas where residents feed nutria during winter months, their population size is gradually growing. This trend is further evidenced by a significant increase in official hunting records in recent years: around 1000 individuals in 2010, more than 5000 in 2015, and a staggering 12 580 in 2021 (CZSO, 2024). However, the actual numbers could be several times higher as a large proportion of hunters do not report their catches.
In the Czech Republic, the nutria was initially considered a non-native species with no apparent impact on nature. However, when overpopulated, it damages aquatic and wetland ecosystems to a significant degree, primarily changing the species composition of plants and reducing green biomass (Grace and Ford, Reference Grace and Ford1996; Evers et al., Reference Evers, Sasser, Gosselink, Fuller and Visser1998; Shaffer et al., Reference Shaffer, Day, Hunter, Lane, Lundberg, Wood and Kandalepas2015). With its invasion, nutria can push native species out of the landscape. It also brings the risk of introducing diseases such as tularaemia, brucellosis, leptospirosis or toxoplasmosis to domestic species (Scheuring, Reference Scheuring1990; Howerth et al., Reference Howerth, Reeves, McElveen and Austin1994; Bounds et al., Reference Bounds, Sherfy, Mollett, Feldhamer, Thompson and Chapman2003; Martino et al., Reference Martino, Stanchi, Silvestrini, Brihuega, Samartino and Parrado2014).
Parasites introduced alongside their hosts into novel territories can profoundly alter natural host–parasite dynamics, thus posing health hazards to native biota and gradually influencing the demography of sympatrically coexisting native fauna (Prenter et al., Reference Prenter, MacNeil, Dick and Dunn2004; Dunn, Reference Dunn2009; Britton, Reference Britton2012). The nutria is host to a relatively high number of parasitic species, with some of them having a zoonotic potential. In its native range, the nutria can harbour more than 30 different helminth species (see Martino et al., Reference Martino, Radman, Parrado, Bautista, Cisterna, Silvestrini and Corba2012; Fugassa, Reference Fugassa2020). Although the helminth communities in introduced areas are slightly altered, the majority of the parasitic taxa overlap with those from native range populations. However, some parasitic species were recorded only in the non-native range [e.g. Trichostrongylus duretteae (Zanzani et al., Reference Zanzani, Di Cerbo, Gazzonis, Epis, Invernizzi, Tagliabue and Manfredi2016) and Taenia taeniaeformis (Umhang et al., Reference Umhang, Richomme, Boucher, Guedon and Boué2013)].
One representative of zoonotic species, which was previously recorded only within non-native nutria populations, is Echinococcus multilocularis [i.e. in France, Germany and Slovenia (Oksanen et al., Reference Oksanen, Siles-Lucas, Karamon, Possenti, Conraths, Romig, Wysocko, Mannocci, Mipatrini, La Torre, Boufana and Casulli2016; Romig and Wassermann, Reference Romig and Wassermann2024)]. Although information on echinococcosis (a disease caused by Echinococcus tapeworms) in nutria is limited, a study in western Germany found that feral nutrias are susceptible to E. multilocularis infection, but less so than muskrats from the same habitat (Hartel et al., Reference Hartel, Spittler, Doering, Winkelmann, Hoerauf and Reiter-Owona2004). In a French zoo, a captive-born nutria was found to have echinococcosis, presumably introduced through the feces of free-roaming foxes (Umhang et al., Reference Umhang, Richomme, Boucher, Guedon and Boué2013). Even though this species was not previously recorded from nutria in the Czech Republic, it is a common parasite of foxes in the region, which serve as definitive hosts, and in some counties the prevalence among free roaming foxes reaches more than 50% (i.e. Kolářová et al., Reference Kolářová, Pavlásek and Chalupský1996; Pavlásek Reference Pavlásek1998; Martínek et al., Reference Martínek, Kolářová and Červený2001, and data provided by the State Veterinary Administration of the Czech Republic from their previous screenings in 2011). The other species with zoonotic potential is Strongyloides myopotami, a specialist of the nutria that causes cutaneous infections in humans known as ‘nutria itch’. Although they do not cause true strongyloidiasis, repeated exposure to larvae of this species may cause an outbreak of severe dermatitis (Little, Reference Little1965). This species was previously documented in nutria in the Czech Republic only among farm-bred animals (Nechybová et al., Reference Nechybová, Langrová and Tůmová2018).
Information on the parasites of nutria in the non-native range (i.e. Europe) is still rather scarce. Previous studies suggest that the species diversity in this range is lower in comparison with the native range (e.g. Lewis and Ball, Reference Lewis and Ball1984; Nardoni et al., Reference Nardoni, Angelici, Mugnaini and Mancianti2011; Umhang et al., Reference Umhang, Richomme, Boucher, Guedon and Boué2013; Zanzani et al., Reference Zanzani, Di Cerbo, Gazzonis, Epis, Invernizzi, Tagliabue and Manfredi2016; Kellnerová et al., Reference Kellnerová, Holubová, Jandová, Vejčík, McEvoy, Sak and Kváč2017; Nechybová et al., Reference Nechybová, Langrová and Tůmová2018; Ježková et al., Reference Ježková, Limpouchová, Prediger, Holubová, Sak, Konečný, Květoňová, Hlásková, Rost, McEvoy, Rajský, Feng and Kváč2021). While the nutria has retained some of the specialist species from its native range [e.g. Trichuris myocastoris (Rylková et al., Reference Rylková, Tůmová, Brožová, Jankovská, Vadlejch, Čadková, Frýdlová, Peřinková, Langrová, Chodová, Nechybová and Scháňková2015) or S. myopotami (Zanzani et al., Reference Zanzani, Di Cerbo, Gazzonis, Epis, Invernizzi, Tagliabue and Manfredi2016)], it has also acquired some new, often generalist, parasite taxa in non-native areas [e.g. Trichostrongylus durettae (Zanzani et al., Reference Zanzani, Di Cerbo, Gazzonis, Epis, Invernizzi, Tagliabue and Manfredi2016) or E. multilocularis (Romig and Wassermann, Reference Romig and Wassermann2024)]. Nonetheless, there is little comprehensive information on the parasites of free-ranging nutria in the Czech Republic (i.e. Nechybová et al., Reference Nechybová, Langrová and Tůmová2018; Ježková et al., Reference Ježková, Limpouchová, Prediger, Holubová, Sak, Konečný, Květoňová, Hlásková, Rost, McEvoy, Rajský, Feng and Kváč2021). In order to compare the diversity of helminth species harboured by Czech nutrias with populations in other countries, the aims of the present study were: (1) to collect data on helminth parasites from different nutria populations in the Czech Republic, (2) to compare the compositions of parasite communities between these populations, and lastly (3) to acquire molecular data for each of the collected helminth species to elucidate genetic structure and intraspecific variability among populations of parasites.
Material and methods
Study area and specimen collection
From January to March 2022, a total of 46 nutria were collected at 8 locations within the Morava catchment in the Czech Republic (Fig. 1, Table 1). The retrieved individuals were placed in plastic bags, and their location, sex and weight were recorded. After transportation to the necropsy room, the viscera were excised and stored frozen.

Figure 1. Locations of the sites within the Czech Republic where the investigated nutria individuals were collected. (1) Šumperk; (2) Ústí nad Orlicí; (3) Olomouc; (4) Brno-city; (5, 6) Brno-vicinity; (7) Hodonín; (8) Břeclav.
Table 1. List of collection sites and number of processed nutria individuals
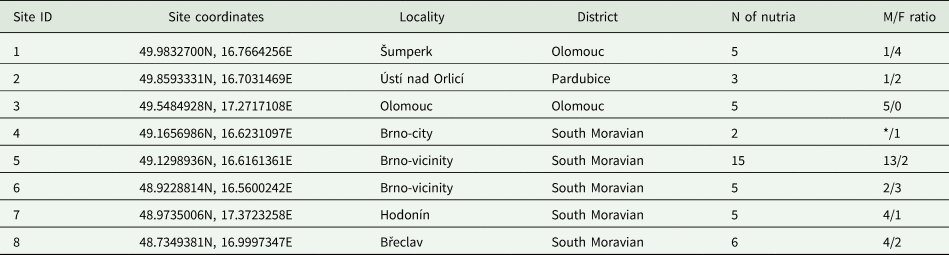
An asterisk at the sex ratio represents the unidentifiable sex of the specimen.
Parasite sampling, fixation and identification
Before parasitological examination, the viscera were gradually thawed. The lungs and liver were examined as squash preparations under an SZX7 stereomicroscope (Olympus, Japan). The heart was cut open using scissors and examined macroscopically. The stomach, small intestine, caecum and colon were longitudinally incised, and the contents of the individual organs were separately diluted with water, homogenized and cleaned using a sieve system with the smallest mesh size of 150 μm. Subsequently, the material retained on the sieve, as well as the walls of the individual organs, was examined under the stereomicroscope.
The collected helminths were counted and stored in 70% ethanol for further morphological analyses. When possible, at least 5 individuals of each helminth species per host specimen were preserved in 96% pure-grade ethanol for DNA extraction. Before fixation in Canada balsam, digeneans were stained in iron acetocarmine, following the protocol of Georgiev et al. (Reference Georgiev, Biserkov and Genov1986). The species identification of digeneans and selection of measurements for morphometric comparison followed the procedures outlined by Gibson et al. (Reference Gibson, Jones and Bray2002), Jones et al. (Reference Jones, Bray and Gibson2005) and Bray et al. (Reference Bray, Gibson and Jones2008). Nematodes were mounted on slides, covered in a mixture of glycerine and water (in a ratio of 3:7), and cleared by gradually increasing the volume of glycerol, according to Moravec (Reference Moravec2013).
To support the findings from the parasitological necropsy, fresh fecal samples available from 20 out of all examined individuals were subjected to coproscopical examination using the modified Sheather's sugar flotation method (Sheather, Reference Sheather1923; Jirků-Pomajbíková and Hůzová, Reference Jirků-Pomajbíková, Hůzová, Modrý, Pafčo, Petrželková and Hasegawa2018). Briefly, walnut-sized fecal samples were homogenised with water using a mortar and pestle, sieved into a tube and centrifuged at 2000 rpm for 3 min. The supernatant was removed, and the sediment was mixed with a sugar solution with a density of 1.3 g/cm3. The sample was centrifuged again at 2000 rpm for 3 min. A surface film containing parasite stages was transferred to a slide with an inoculating loop, covered with a coverslip and examined under a light microscope (Foreyt, Reference Foreyt2002).
Primary epidemiological data, including prevalence, mean abundance and minimum and maximum intensities of infection, were calculated for each parasite species according to Bush et al. (Reference Bush, Lafferty, Lotzs and Shostakll1997). Prevalence was defined as the percentage of host individuals infected by a given parasite species, and mean abundance was calculated as the mean number of parasite specimens per individual host considering both infected and uninfected hosts. Following the suggestion of Rózsa et al. (Reference Rózsa, Reiczigel and Majoros2000) for interpreting epidemiological data, a confidence interval at the level of 95% was calculated for mean abundance.
PCR amplification, sequence analysis and phylogenetic analyses
Total DNA from fresh or frozen fecal samples (N = 40) was isolated using PowerSoil DNA isolation kit (Qiagen Company, USA). The extracted fecal DNA was screened by qPCR for the detection of Strongyloides (18S rDNA) using a Real-Time PCR LightCycler® 480 (Roche, Switzerland). The primers and reaction conditions used followed Verweij et al. (Reference Verweij, Canales, Polman, Ziem, Brienen, Polderman and van Lieshout2009).
Parasite genomic DNA was extracted using NucleoSpin® Tissue kit (Macherey-Nagel, Düren, Germany) following the manufacturer's protocol. Prior to extraction, the specimens (or their parts) were removed from the ethanol and dried in a thermal block. The PCR reactions were performed in a 20 μL reaction mixture containing 14 μL nuclease-free water, 4 μL FIREPol Master Mix Ready to Load (Solis BioDyne, Tartu, Estonia), 0.5 μm of each primer and 1 μL of DNA template. PCR products were detected by electrophoresis in 1% agarose gels stained with GoodView (SBS Genetech, Bratislava, Slovakia). A total of 8 different primer combinations were used for the amplification of specific genomic regions in nematodes, trematodes and cestodes. A list of primer sequences and cycling conditions is provided in Table 2. The resulting amplicons were subsequently subjected to Sanger sequencing, after which the obtained sequences were compared in silico with data available in publicly accessible databases and were also used for phylogenetic analyses.
Table 2. List of primers used for PCR amplification of mitochondrial and nuclear markers in the present study
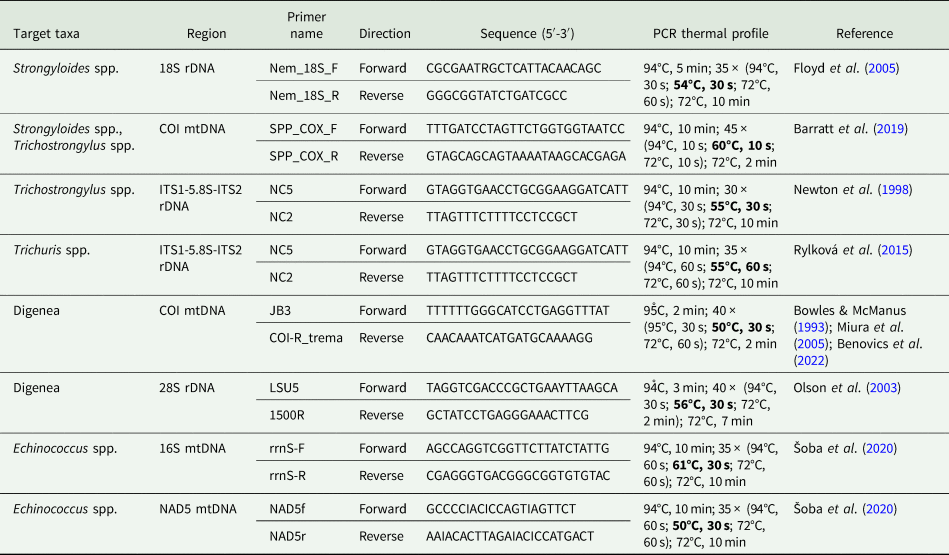
In order to assess the phylogenetic position of selected parasite taxa (i.e. in the case of strongylids) and/or selected haplotypes (i.e. in the case of COI variants of various Strongyloides spp.), additional orthologous sequences from congeners or different phylogenetically close species were obtained from GenBank (accession numbers are included within phylogenetic trees). The sequences were aligned by means of the fast Fourier transform algorithm implemented in MAFFT (Katoh et al., Reference Katoh, Misawa, Kuma and Miyata2002), using the G-INS-i refinement method. A general time-reversible model (GTR; Lanave et al., Reference Lanave, Preparata, Saccone and Serio1984) was applied for each partition of the alignment, each representing an individual genomic region. For the Strongyloides spp. dataset, the GTR model was applied for the entire length of the alignment. Phylogenetic trees were constructed using Bayesian inference (BI) and maximum likelihood (ML) approaches in MrBayes 3.2. (Ronquist et al., Reference Ronquist, Teslenko, van der Mark, Ayres, Darling, Höhna, Larget, Liu, Suchard and Huelsenbeck2012) and RAxML 8.1.12 (Stamatakis, Reference Stamatakis2006, Reference Stamatakis2014), respectively. The BI analysis used the Metropolis-coupled Markov chain Monte Carlo algorithm with 2 parallel runs of 1 cold and 3 hot chains, and was run for 106 generations, sampling trees every 100 generations. The initial 30% of all saved trees were discarded as ‘burn-in’ after checking that the standard deviation split frequency fell below 0.01. The convergence of the runs and the parameters of individual runs were checked using Tracer v. 1.7.1 (Rambaut et al., Reference Rambaut, Drummon, Xie, Baele and Suchard2018). Posterior probabilities for each tree node were calculated as the frequency of samples recovering a given clade. The clade bootstrap support for ML trees was assessed by simulating 103 pseudoreplicates.
Results
Nutria's helminth diversity and epidemiology
A total of 6 helminth species were collected from 46 nutria specimens during microscopical examination (Table 3). The highest prevalence (67.4%) was recorded for S. myopotami, which parasitized nutrias from all collection sites (Fig. 2). This species also exhibited the highest mean abundance and maximum intensity of infection among the examined individuals. The flotation method revealed this species (as infectious eggs) in each fresh fecal sample (Fig. 3), even when the examination of the viscera did not yield any adult specimens. After combining the data from necropsy and flotation, the overall prevalence increased to 78.3%. In addition, qPCR analysis confirmed the presence of Strongyloides DNA in 100.0% of the tested fecal samples, further corroborating its high prevalence. Combining all 3 methods, the total prevalence of S. myopotami reached 93.5% among all processed nutria individuals. The second most prevalent species was T. myocastoris, which was, during dissection, recorded in nutria from all sites except Olomouc (site 3) and Břeclav (site 8). Molecular analysis revealed that this species was genetically identical in the ITS1-5.8S-ITS2 region to T. myocastoris specimens previously documented in the Czech Republic (MF077367). Echinococcus multilocularis hydatids (Fig. 4) were found in the livers of 5 individuals, altogether from 3 sites (Table 3). A further unidentified trematode species was also found in 5 nutria individuals (collected from 2 nutrias from site 6, 2 nutrias from site 8 and 1 nutria from site 3). Due to the poor quality of the material, especially as a result of deep freezing the cadavers, morphological identification of these trematodes was not possible. However, the molecular sequences (partial 28S rDNA and partial COI mtDNA regions) suggest that these specimens belonged to the family Psilostomidae, as they had 99.8% similarity to a sequence of Psilostomidae gen. sp. in GenBank (MN726950; see Table 3 with accession numbers for the newly obtained DNA sequences). Trichostrongylus nematodes were also recorded in nutria from 3 collection sites, but could not be identified to species level due to the insufficient number and quality of specimens for morphological evaluation (Fig. 5). Additionally, a single nutria individual, collected in the vicinity of Brno city (site 6), was parasitized by Echinostoma trematodes. Only 2 specimens of this helminth species were collected and all visible morphological features resembled Echinostoma revolutum.
Table 3. List of collected parasite species identified during microscopical examination, with localization, basic epidemiological data, sites with positive records, GenBank accession numbers to newly generated sequences and information about previous records from nutria in the Czech Republic
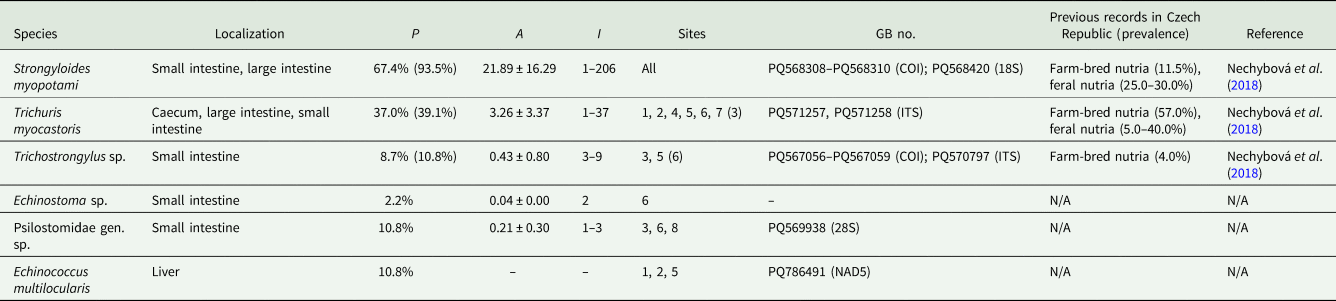
P = prevalence; A = mean abundance with confidence interval at the level of 0.95; I = intensity of infection (min–max); GB no. = GenBank accession numbers to the representative sequences. Prevalence values and site numbers in brackets are for records including flotation and quantitative PCR results. Dashes (–) indicate the species was recorded in an uncountable number of cysts. N/A indicates that this parasite taxon was not previously recorded in nutria from the Czech Republic.
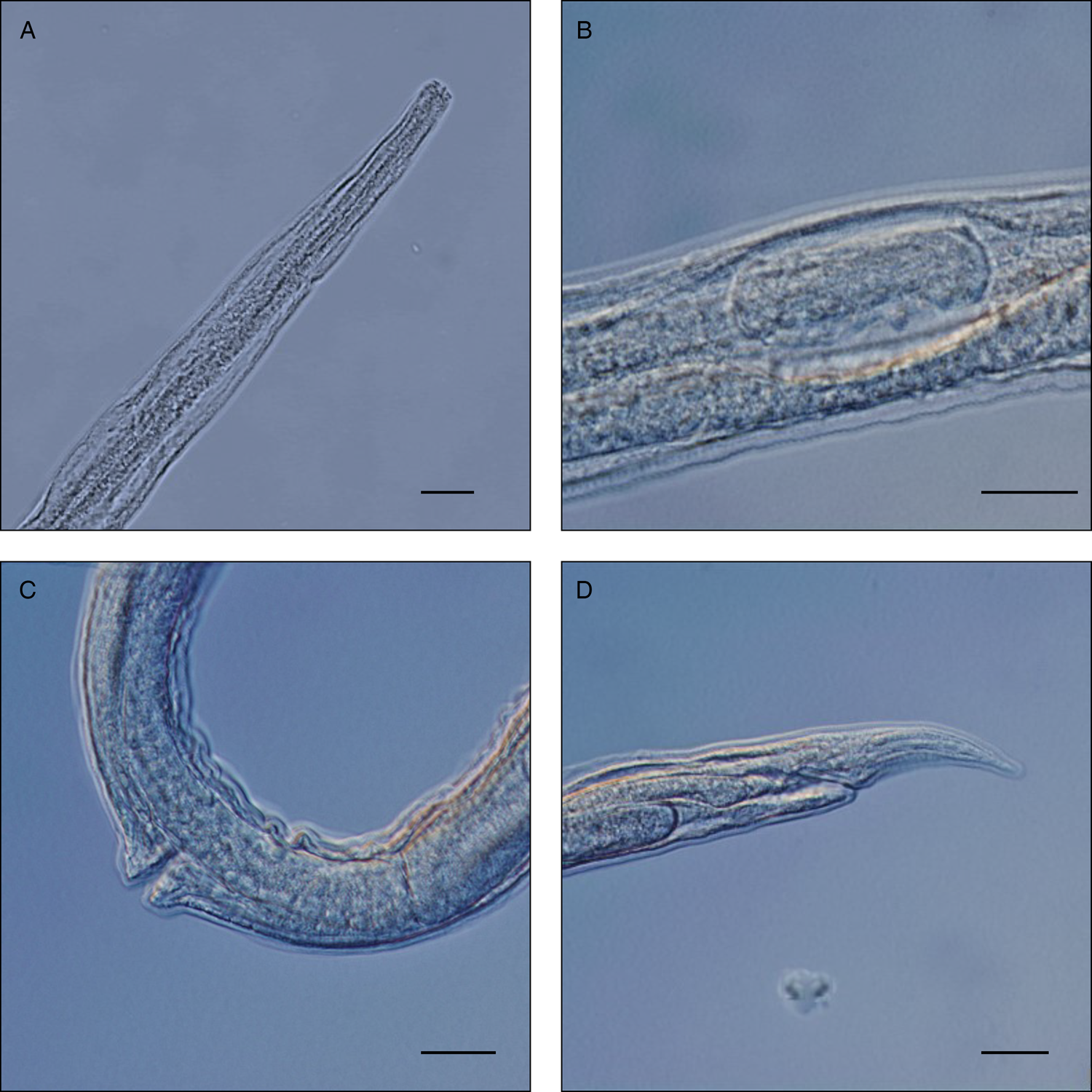
Figure 2. Microscopic details of Strongyloides myopotami parasitic females. Scale bar = 20 μm. (A) anterior end; (B) egg in the uterus; (C) detail of vulva; (D) posterior end with tail and anus.
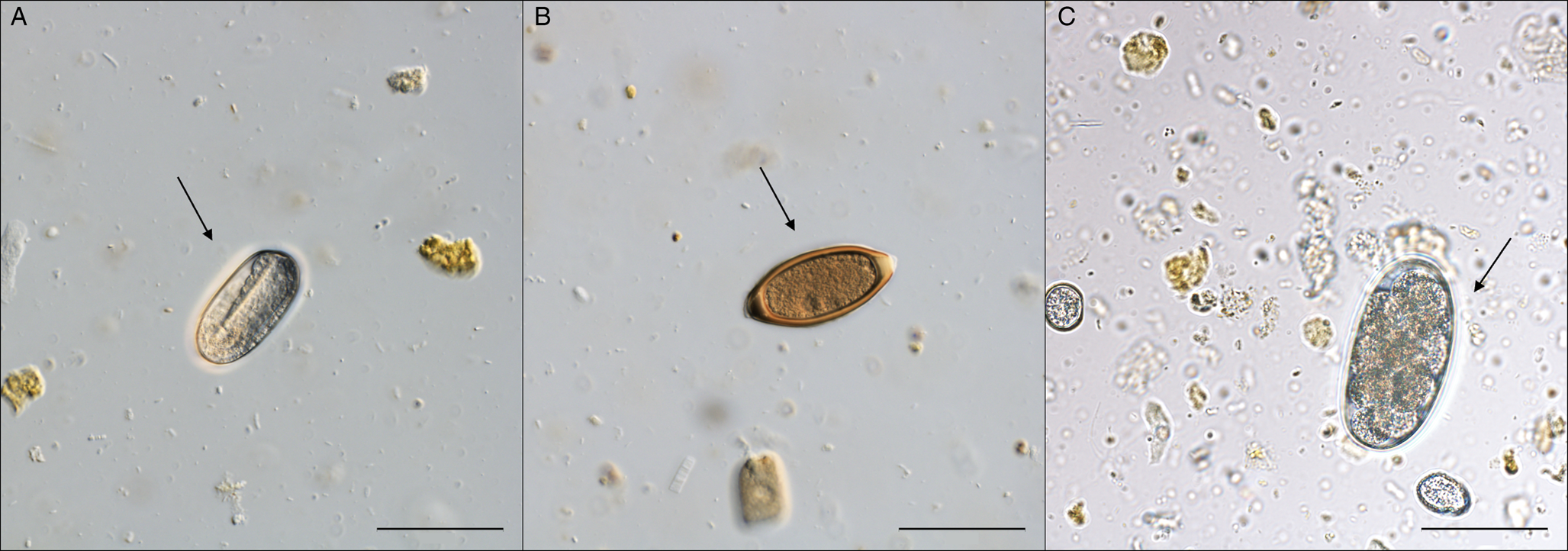
Figure 3. Egg stages of Nematoda detected in nutria fecal samples using Sheather's sugar flotation. Scale bar = 50 μm. (A) Oval egg with so-called U-larva of Strongyloides myopotami; (B) egg of Trichuris myocastoris; (C) egg of strongylid nematode, probably Trichostrongylus sp.
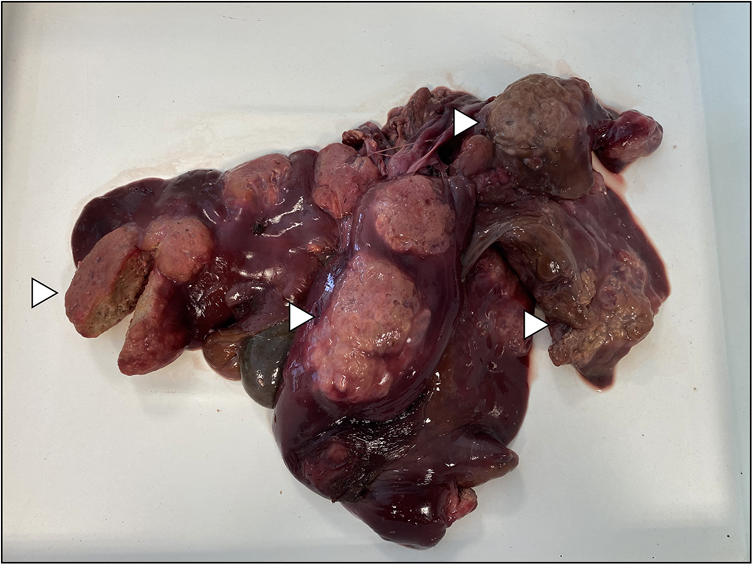
Figure 4. Alveolar hydatid cysts of Echinococcus multilocularis in the liver of nutria collected near the city of Šumperk. Examples of cysts are pinpointed by white arrows.
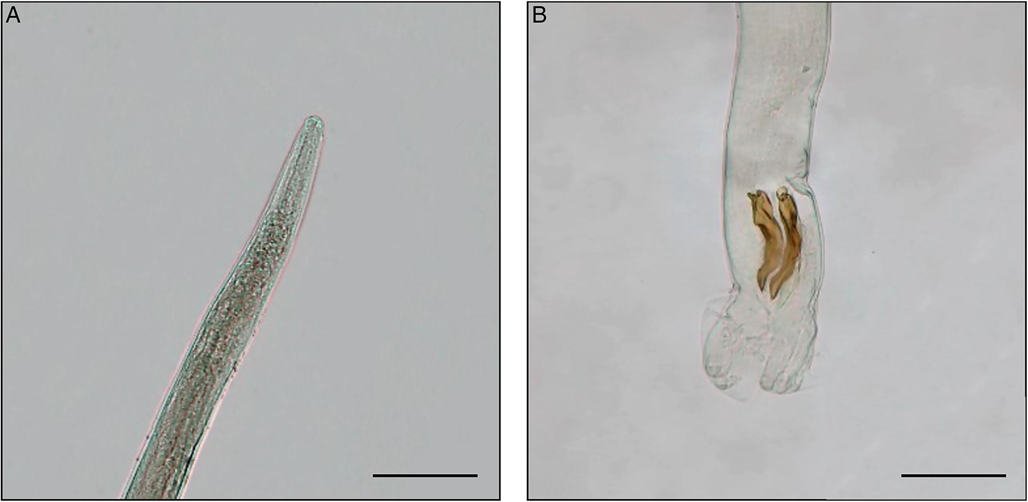
Figure 5. Microscopic details of Trichostrongylus sp. specimen. Scale bar = 100 μm. (A) The anterior end of the male; (B) the posterior end of the male with spicules and copulatory bursa.
Genetic diversity and variability in obtained nematode species
All obtained helminth species were sequenced to analyse genetic diversity and intraspecific variability. For trematodes, no intraspecific variability was observed in either the 28S or COI region. For T. myocastoris and Trichostrongylus sp. only a minor intraspecific variability was observed in COI and no variability in ITS regions. A total of 49 S. myopotami specimens were sequenced, all of which were identical in the 18S region to the S. myopotami sequence deposited in GenBank (AB453313). The amplified COI region spanned 240 nucleotide positions, and 3 distinct genetic variants were identified among these individuals. Haplotype I was observed in individuals at all sites where S. myopotami was present. Haplotype II was absent in individuals from Šumperk (site 1). All 3 haplotypes were found only in individuals from the Brno-vicinity (site 6), with only 1 sequenced individual carrying haplotype III. Haplotypes II and III were the most similar, differentiating in only 1 substitution. Only a single S. myopotami specimen was collected from Ústí nad Orlicí; however, PCR amplification did not yield products of sufficient quality, and therefore, it was not possible to assess the haplotype present at this site.
The resulting Strongyloides phylogenetic tree revealed 12 separate clades corresponding to different Strongyloides species. For the partial COI mtDNA region of Strongyloides, the alignment consisted of 26 sequences, including Necator americanus as the outgroup. Both BI and ML analyses generated trees with identical topologies. The BI tree with posterior probabilities and bootstrap values (corresponding to the ML tree) along respective nodes is presented in Fig. 6. All the haplotypes of S. myopotami from nutrias clustered within a highly supported subclade. However, its position was not fully resolved due to the basal polytomy of the tree.
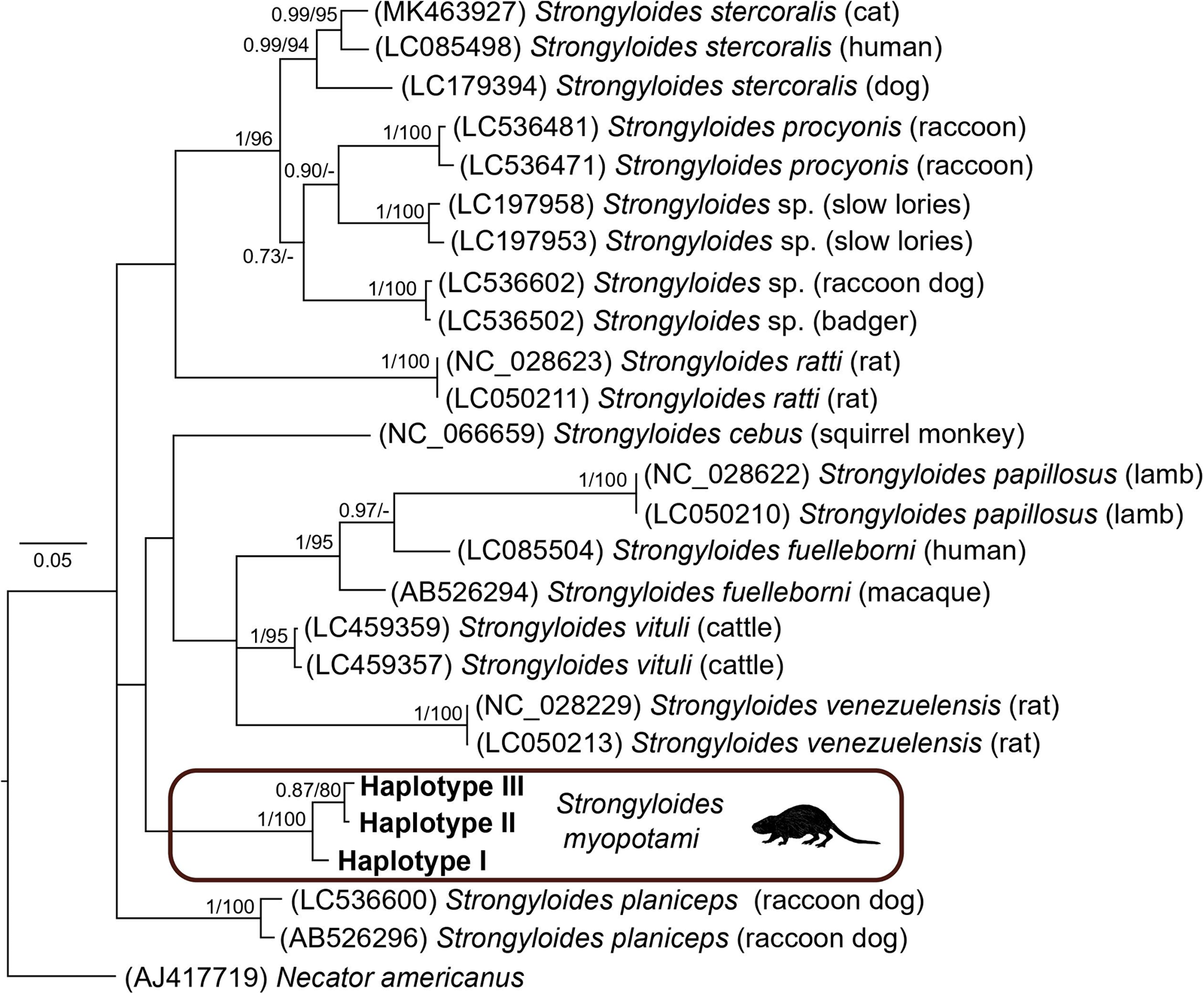
Figure 6. Phylogenetic tree of 25 sequences of Strongyloides species reconstructed by Bayesian inference. The tree is rooted using Necator americanus as an outgroup. Values at the nodes indicate posterior probabilities from BI and bootstrap values from ML analyses. Dashes indicate nodal support values below 0.70 and 50, respectively. The hosts of respective Strongyloides specimens are noted in brackets.
The final sequence alignment, encompassing selected trichostrongylid species and the outgroup Ancylostoma braziliense, was constructed using the ITS1-5.8S-ITS2 region. The alignment included 44 taxa and spanned 775 unambiguously aligned nucleotide positions. The BI tree, with posterior probabilities and bootstrap values (corresponding to the ML tree) along respective nodes, is presented in Fig. 7. Trichostrongylus spp. formed a well-supported monophyletic group, with the sister position to the Libyostrongylus spp. The Trichostrongylus sp. collected from nutria in the Czech Republic clustered within the ‘Trichostrongylus’ clade; however, its position was not fully resolved, resulting in polytomy. This Trichostrongylus sp. shared 98.3% sequence similarity with T. vitrinus which was collected from Roe deer (Capreolus capreolus) in Russia. The species collected from nutrias in the Czech Republic exhibited greater genetic dissimilarity (>2%) to T. axei or T. colubriformis, suggesting it represents a distinct species.
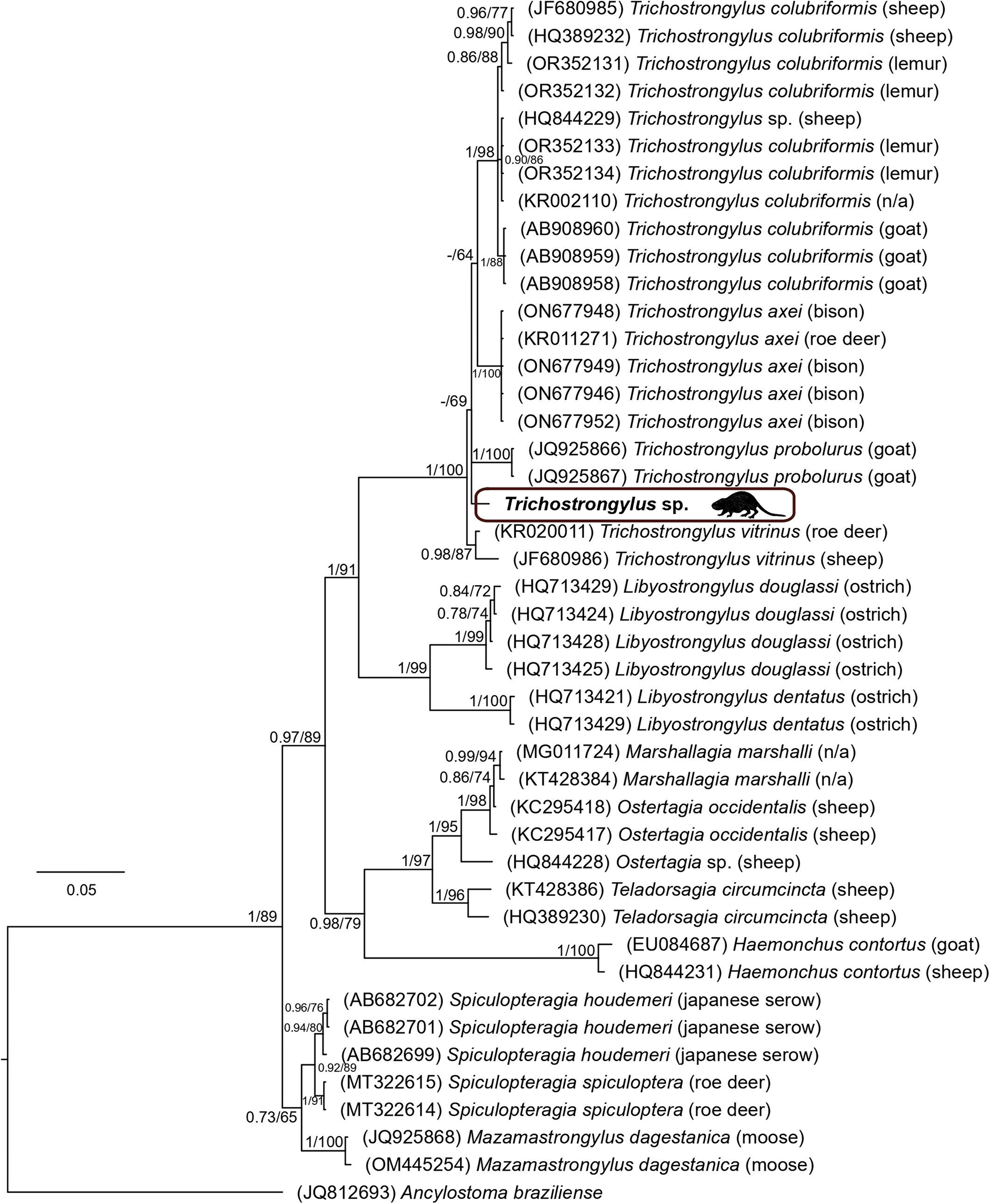
Figure 7. Phylogenetic tree of 43 sequences of 15 trichostrongylid species reconstructed by Bayesian inference. The tree is rooted using Ancylostoma braziliense as an outgroup. Values at the nodes indicate posterior probabilities from BI, and bootstrap values from ML analyses. Dashes indicate nodal support values below 0.70 and 50, respectively. The hosts of respective trichostrongylid specimens are noted in brackets. N/A indicates that the host information is not available.
The sequencing of 2 mitochondrial genomes supported the identification of the hydatid cysts as E. multilocularis. The individuals collected from nutrias in the Czech Republic were genetically identical in the NADH region to those from nutrias in Slovenia (MW560732). In contrast, in the 16S rRNA region, they were identical to specimens from France (e.g. the sequence retrieved from the complete mitochondrial genome; OQ599967) and differed by a single nucleotide substitution from those collected in Slovenia (MW558108).
Discussion
Since the introduction of nutria to the Czech Republic, its population size has been steadily increasing. Despite its established status as an invasive species in the region, the parasite fauna associated with nutria remains critically understudied. In its native range, 64 parasite taxa have been recorded, which were assigned either to species level or at least to a higher taxonomic level (e.g. El-Kouba et al., Reference El-Kouba, Marques, Pilati and Hamann2009; Issia et al., Reference Issia, Pietrokovsky, Sousa-Figueiredo, Stothard and Wisnivesky-Colli2009; Rossin et al., Reference Rossin, Varela and Timi2009; Martino et al., Reference Martino, Radman, Parrado, Bautista, Cisterna, Silvestrini and Corba2012; Benati et al., Reference Benati, Moraes, Hoppe, Tebaldi, Júnior and da Costa Freitaz2017; Fugassa, Reference Fugassa2020). In contrast, European studies have documented only 11 of these parasites in nutria, with an additional 13 taxa identified exclusively in European nutria populations (Lewis and Ball, Reference Lewis and Ball1984; Ménard et al., Reference Ménard, Agoulon, L'Hostis, Rondelaud, Collard and Chauvin2001; Nardoni et al., Reference Nardoni, Angelici, Mugnaini and Mancianti2011; Umhang et al., Reference Umhang, Richomme, Boucher, Guedon and Boué2013; Zanzani et al., Reference Zanzani, Di Cerbo, Gazzonis, Epis, Invernizzi, Tagliabue and Manfredi2016; Nechybová et al., Reference Nechybová, Langrová and Tůmová2018; Ježková et al., Reference Ježková, Limpouchová, Prediger, Holubová, Sak, Konečný, Květoňová, Hlásková, Rost, McEvoy, Rajský, Feng and Kváč2021). Thus far, 24 species have been identified in Europe, indicating that the diversity of nutria parasites is markedly higher in their native range compared to the regions where they have been introduced.
Information on nutria parasites in the Czech Republic is scarce and only a few studies have provided any comprehensive insight, based on a combination of diagnostic methods. Regarding helminths, only 2 genera were previously recorded; S. myopotami was recorded among fur-farmed animals in the Czech Republic with 25% prevalence (out of 20 examined animals, Nechybová et al., Reference Nechybová, Langrová and Tůmová2018). In the same study, also a further unidentified Strongyloides species was detected by coprological methods in free roaming nutrias at various sites in the Czech Republic, although it is highly possible it will also be S. myopotami, in accordance with our study. The other taxon was T. myocastoris, which was recorded among farmed animals (Rylková et al., Reference Rylková, Tůmová, Brožová, Jankovská, Vadlejch, Čadková, Frýdlová, Peřinková, Langrová, Chodová, Nechybová and Scháňková2015; Nechybová et al., Reference Nechybová, Langrová and Tůmová2018) and feral animals, as well (Nechybová et al., Reference Nechybová, Langrová and Tůmová2018).
On the basis of the necropsy of 46 nutrias, the coproscopy of feces from 20 individuals, the amplification of 40 extracted fecal DNA samples using qPCR (18S rDNA) and sequencing of the 18S ribosomal subunit DNA and partial COI mtDNA, the highest recorded prevalence was of S. myopotami (93.5%). When coproscopy and molecular analyses were not considered, the prevalence of this parasite appeared lower. Specifically, the flotation method revealed the presence of this parasite in 5 nutrias, although no adult parasites were found during the necropsy of those individuals. This discrepancy suggests that the infection intensity in these cases may have been too low to be detected during necropsy. In contrast, qPCR analysis, the most sensitive method employed, detected Strongyloides in 100% of the analysed fecal samples. Previous studies have also reported high prevalence rates of S. myopotami. Choe et al. (Reference Choe, Lee, Park, Oh, Jeon and Eom2014) documented a 100.0% prevalence in a survey of 10 nutrias in Korea. Babero and Lee (Reference Babero and Lee1961) recorded a prevalence of 62.5% in 56 nutrias from Louisiana (USA). In the native range of nutria, a prevalence of 26.7% was observed (Martino et al., Reference Martino, Radman, Parrado, Bautista, Cisterna, Silvestrini and Corba2012). The highest prevalence in Europe to date was found in Italy, at 63.4% (Zanzani et al., Reference Zanzani, Di Cerbo, Gazzonis, Epis, Invernizzi, Tagliabue and Manfredi2016). Our current study also ranks among those in which S. myopotami was the most commonly represented parasite, and its prevalence among Czech nutria populations appears to be comparatively higher, as previously reported by Nechybová et al. (Reference Nechybová, Langrová and Tůmová2018) (25–30%).
The second most common parasite was T. myocastoris (prevalence 37.0%). Its significance as a common parasite of nutrias is corroborated by Babero and Lee (Reference Babero and Lee1961), who recorded prevalences of 28 and 50% at 2 study sites in Louisiana (USA). Similarly, Martino et al. (Reference Martino, Radman, Parrado, Bautista, Cisterna, Silvestrini and Corba2012) observed T. myocastoris among the most numerous parasite species in South America, with a prevalence of 13.8%. In the Czech Republic, Nechybová et al. (Reference Nechybová, Langrová and Tůmová2018) recorded a prevalence of 40% for T. myocastoris in a study of wild nutrias and a prevalence of 5% for Trichuris sp. in fecal samples from farmed nutrias.
The genus Trichostrongylus could not be definitively identified at the species level. Identification is primarily based on the shape and size of male spicules and copulatory bursa, while females are consistently difficult to identify. According to Dikmans’ (Reference Dikmans1937) key, the Trichostrongylus specimens in our study most closely resembled T. ransomi in spicule shape and length. Another distinguishing characteristic was the distance of the anus from the tail tip in females. When compared to the work of Ghasemikhah et al. (Reference Ghasemikhah, Mirhendi, Kia, Mowlavi, Sarmadian, Meshgi, Golestan and Mobedi2011), the spicule shape (resembling a high-heeled shoe) was similar to that in T. colubriformis species. Previous studies have reported the presence of Trichostrongylus duretteae, T. sigmodontis, T. colubriformis and T. retortaeformis in nutrias (Babero and Lee, Reference Babero and Lee1961; Zanzani et al., Reference Zanzani, Di Cerbo, Gazzonis, Epis, Invernizzi, Tagliabue and Manfredi2016; Fugassa, Reference Fugassa2020). Based on newly generated DNA sequences, our species was genetically most similar to T. axei. However, sequences for T. sigmodontis or T. duretteae are not available in GenBank, so genetic similarity to these species could not be assessed. Our mitochondrial DNA sequences, representing a more variable region, make comparisons with GenBank data less meaningful unless sequences from the same or closely related organisms are available.
The trematode species with a prevalence of 11% was classified within the family Psilostomidae based on the DNA sequence similarity of the large ribosomal subunit (28S). Mitochondrial DNA (COI) did not assist in identifying the parasite, as no closely related sequence was available in the GenBank database. The family Psilostomidae comprises 13 genera, which are gastrointestinal parasites of birds and mammals. Morphologically, the members of this family resemble those of the family Echinostomatidae, but Psilostomidae lack the characteristic spined collar (Kostadinova, Reference Kostadinova, Jones, Bray and Gibson2005; Atopkin, Reference Atopkin2011). Given the presence of the genus Psilotrema in Europe (Atopkin, Reference Atopkin2011) and the closest genetic similarity observed, it is plausible to hypothesize that our specimens belong to this genus. Additionally, Psilostomum sp. has also been previously reported in nutrias, although only in the metacercarial stage; however, it was not identified in our study (Babero and Lee, Reference Babero and Lee1961).
The presence of E. multilocularis in the native range of nutria remains undocumented. In Europe, the occurrence and increasing incidence of infections with the zoonotic tapeworm E. multilocularis are closely linked with the rising populations of foxes, raccoon dogs and nutrias (Janovsky et al., Reference Janovsky, Bacciarini, Sager, Grone and Gottstein2002; Romig, Reference Romig2009; Križman et al., Reference Križman, Švara, Šoba and Rataj2022). Similarly, the spread of this parasite correlates with growing urban fox populations and the movements of infected dogs (Goodfellow et al., Reference Goodfellow, Shaw and Morgan2006). Within the Morava basin, the presence of echinococcosis was detected in nutria at the Šumperk (site 1), Ústí nad Orlicí (site 2) and Brno-vicinity (5) locations, with the former 2 being only about 20 km apart. Although E. multilocularis was previously recorded among foxes in all 3 respective Czech districts [ranging from 10.0 to 40.0% prevalence, see Kolářová et al. (Reference Kolářová, Matějů, Hozáková, Stejskal, Hrdý, Kolářová, Leissová, Skála and Dundr2017) and references therein], this finding substantiates the Eurasian and somewhat disjunct distribution of the parasite (McManus et al., Reference McManus, Zhang, Li and Bartley2003; Oksanen et al., Reference Oksanen, Siles-Lucas, Karamon, Possenti, Conraths, Romig, Wysocko, Mannocci, Mipatrini, La Torre, Boufana and Casulli2016). Our samples showed a high genetic similarity with E. multilocularis from various European countries, particularly Slovenia and France. This genetic congruence classifies the Moravian nutria specimens within the widely distributed genotype observed across western, central and eastern Europe (Santoro et al., Reference Santoro, Santolamazza, Cacciò, La Rosa, Antolová, Auer, Bagrade, Bandelj, Basso, Beck, Citterio, Davidson, Deksne, Frey, Fuglei, Glawischnig, Gottstein, Harna, Petersen and Karamon2024).
Studies on E. multilocularis tend to focus on the occurrence of the parasite in definitive hosts, particularly foxes. The increase in fox populations in western European countries, attributed to rabies vaccination programmes (Eckert et al., Reference Eckert, Conraths and Rackmann2000), probably prompted the migration of young foxes from areas with high population density to those with lower density, moving eastward (Sréter et al., Reference Sréter, Széll, Egyed and Varga2003). In the 1990s, changes in land use occurred in the former communist countries. Additionally, a decline in fox hunting, driven by a fall in the value of fox fur and the implementation of rabies vaccination programmes in central and eastern European countries, contributed to a rise in the fox population (Szemethy et al., Reference Szemethy, Heltai and Biró2000). This increase in fox numbers likely led to a higher prevalence and spread of E. multilocularis (Sréter et al., Reference Sréter, Széll, Egyed and Varga2003).
Currently, E. multilocularis is widely distributed in Europe (ESFA, 2021). The emergence of a new potential definitive host, the invasive raccoon dog (Sutor, Reference Sutor2008), has also facilitated this spread. This species is now well established in Europe and continues to expand its range westward and southward across the continent (Genovesi, Reference Genovesi2009). However, the prevalence of E. multilocularis in raccoon dogs is lower compared to foxes, which are still considered the primary definitive host for this parasite.
A study conducted by Oksanen et al. (Reference Oksanen, Siles-Lucas, Karamon, Possenti, Conraths, Romig, Wysocko, Mannocci, Mipatrini, La Torre, Boufana and Casulli2016) demonstrated that nutrias generally play a minimal or no role in the life cycle of E. multilocularis within European Union countries. Studies investigating the presence of E. multilocularis in European nutrias found a prevalence of 0.4% from 12 locations in western France (Umhang et al., Reference Umhang, Richomme, Boucher, Guedon and Boué2013) and 5.9% in 2 distinct areas southwest of North Rhine-Westphalia in Germany (Hartel et al., Reference Hartel, Spittler, Doering, Winkelmann, Hoerauf and Reiter-Owona2004). Nonetheless, in areas with moderate to high infection prevalence in red foxes, nutrias may contribute to the life cycle of this parasite (Oksanen et al., Reference Oksanen, Siles-Lucas, Karamon, Possenti, Conraths, Romig, Wysocko, Mannocci, Mipatrini, La Torre, Boufana and Casulli2016). Studies conducted in France and Slovenia have identified nutrias as bioindicators of E. multilocularis presence in the environment (Umhang et al., Reference Umhang, Richomme, Boucher, Guedon and Boué2013; Križman et al., Reference Križman, Švara, Šoba and Rataj2022). Consequently, it is plausible that foxes near the Šumperk, Ústí nad Orlicí and Brno-vicinity locations could also be infected with E. multilocularis.
Data availability statement
The data supporting the conclusions of this study are included in this article. The newly generated sequences were submitted to the GenBank database under accession numbers PQ567056–PQ567059; PQ568308–PQ568310; PQ568420; PQ569938; PQ570797; PQ571257–PQ571258; PQ786491.
Acknowledgements
We are grateful to Lucie Seidlová and Viktória Vanerková for their help during dissections and parasites collection. We also thank Matthew Nicholls for English corrections and proof-reading.
Author contributions
M. B. and O. M. conceived and designed the study. O. M., J. D. and J. S. obtained nutrias examined within this study. M. B., E. N., A. K., L. Š., E. J. and O. M. collected and identified parasitological material. M. B. and A. K. performed morphological analyses; M. B., E. N., A. K. and E. J. performed molecular laboratory procedures; and M. B. and E. N. performed phylogenetic and statistical analyses. M. B. put together the results and wrote the draft together with E. N., L. Š. and O. M. All the other authors revised the draft and approved the final version.
Financial support
This research was funded by the Internal Grant Agency of Mendel University in Brno under grant No. AF-IGA2022-IP-030, titled ‘Quality of parameters of meat of nutria (Myocastor coypus) and its technological evaluation in meat products’. This project was also partially supported by Masaryk University internal grant No. MUNI/A/1602/2023.
Competing interests
None.
Ethical standards
All applicable institutional, and national guidelines for the care and use of invasive animals were followed.













